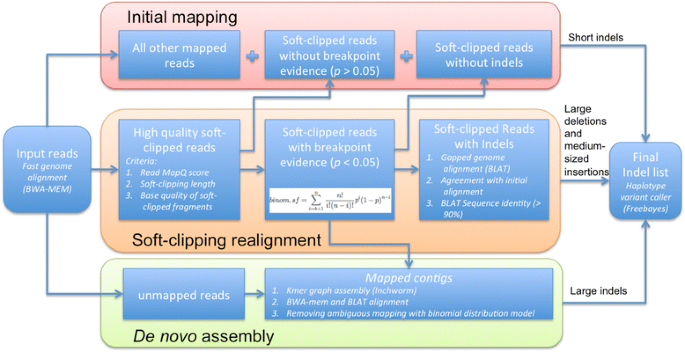
ScanIndel: a hybrid framework for indel detection via gapped alignment, split reads and de novo assembly | Genome Medicine | Full Text

VarDict: a novel and versatile variant caller for next-generation sequencing in cancer research. - Abstract - Europe PMC

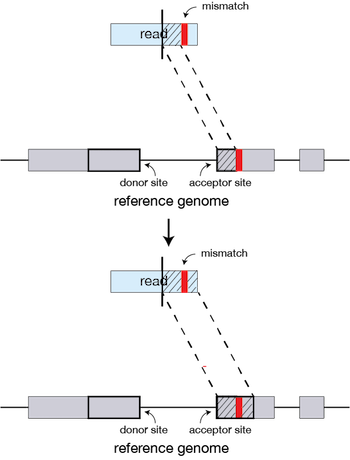
Soft-clipped reads vs. improperly aligned reads at a structural variant... | Download Scientific Diagram
![PDF] Socrates: identification of genomic rearrangements in tumour genomes by re-aligning soft clipped reads | Semantic Scholar PDF] Socrates: identification of genomic rearrangements in tumour genomes by re-aligning soft clipped reads | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/16b34a01c549bc1b12b6c5a17e85e1db117f26e3/2-Figure1-1.png)
PDF] Socrates: identification of genomic rearrangements in tumour genomes by re-aligning soft clipped reads | Semantic Scholar
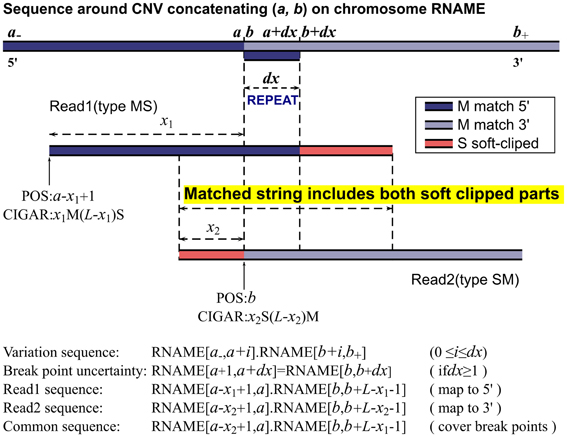
Frontiers | MATCHCLIP: locate precise breakpoints for copy number variation using CIGAR string by matching soft clipped reads
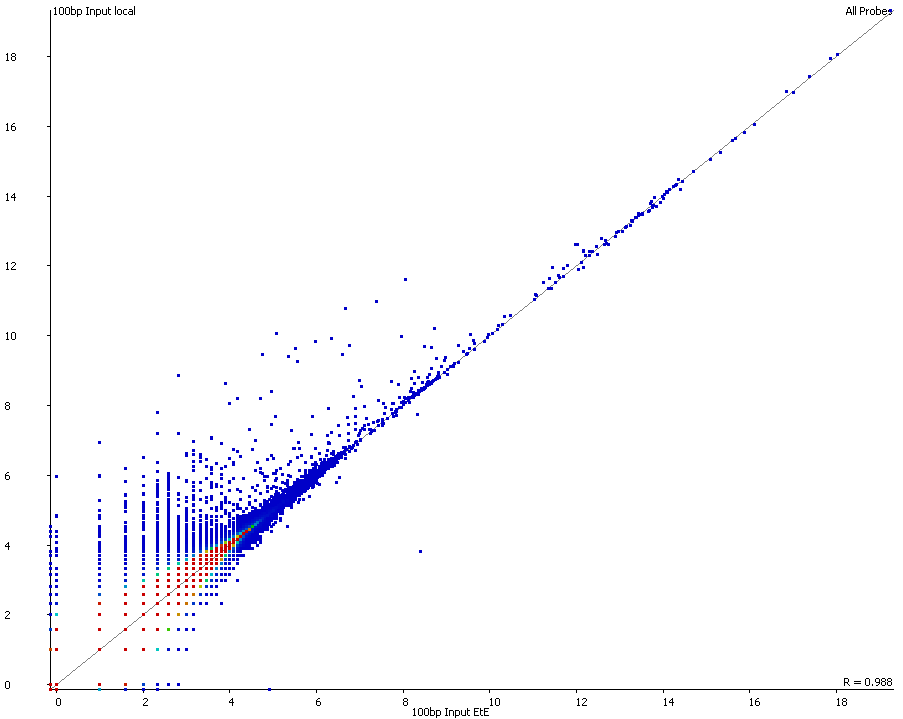
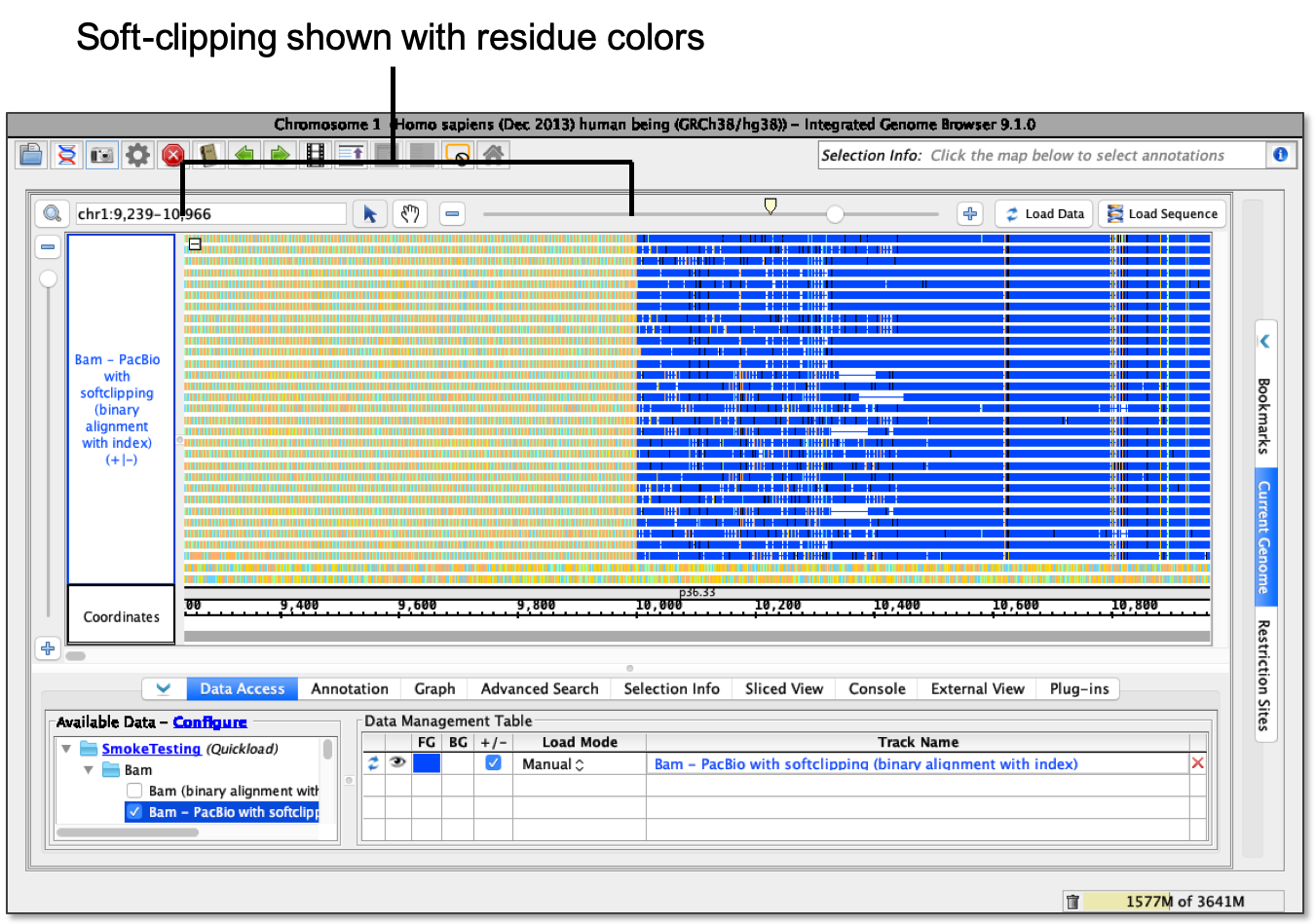
QC Fail Sequencing » Soft-clipping of reads may add potentially unwanted alignments to repetitive regions

eKLIPse: a sensitive tool for the detection and quantification of mitochondrial DNA deletions from next-generation sequencing data - ScienceDirect
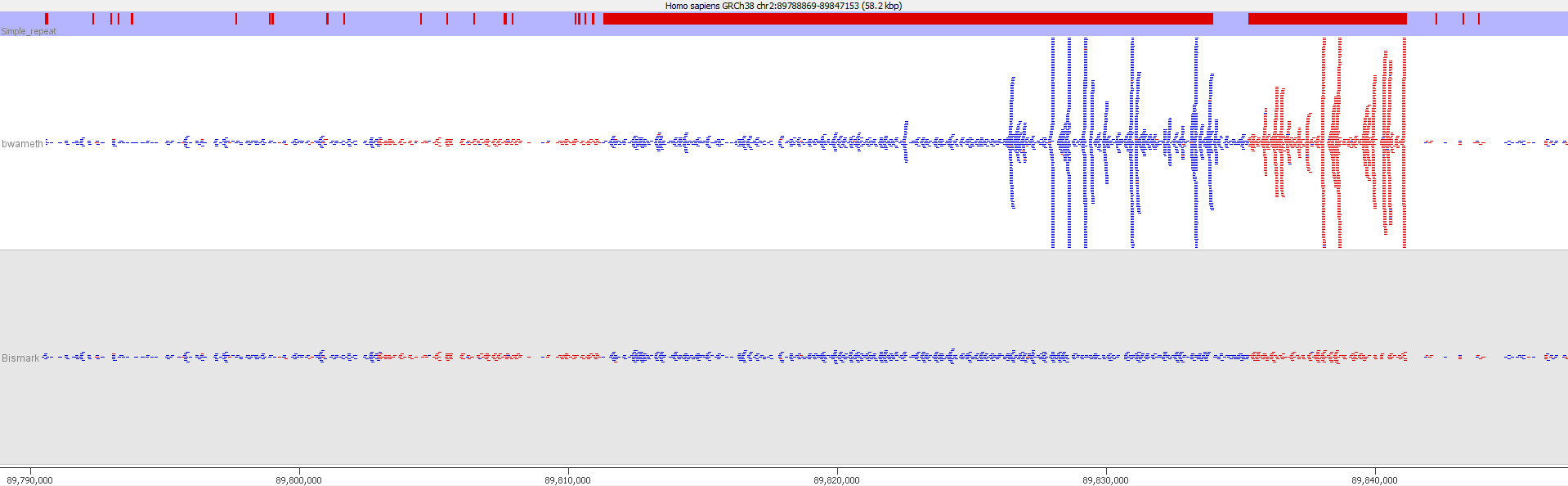
QC Fail Sequencing » Soft-clipping of reads may add potentially unwanted alignments to repetitive regions
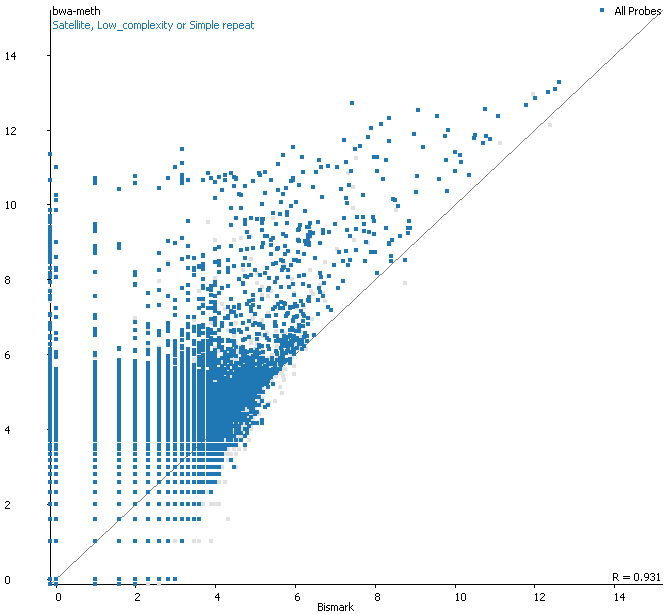
QC Fail Sequencing » Soft-clipping of reads may add potentially unwanted alignments to repetitive regions
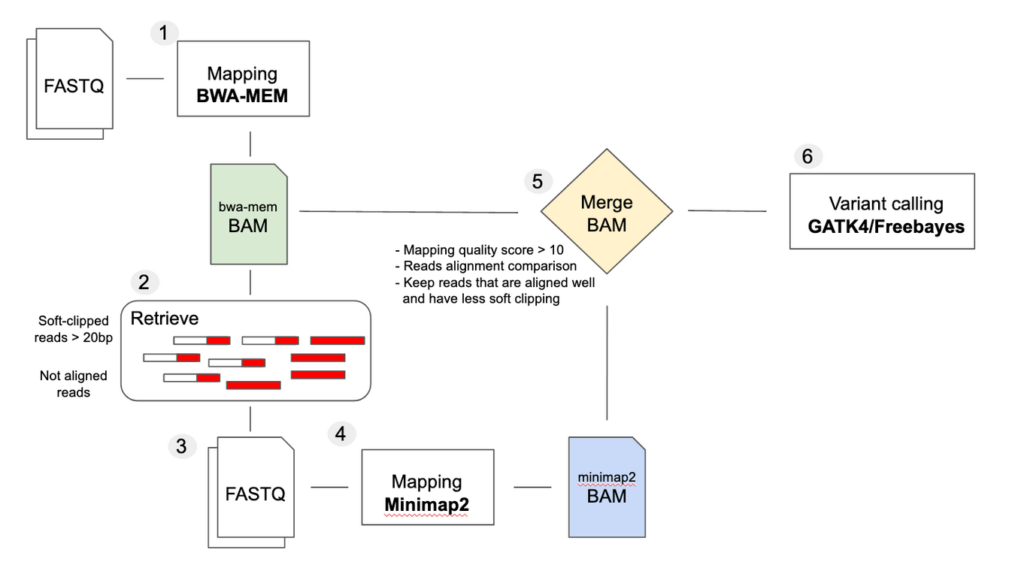
Challenges and importance of mid-sized deletion identification for genomic medicine - SeqOne Genomics
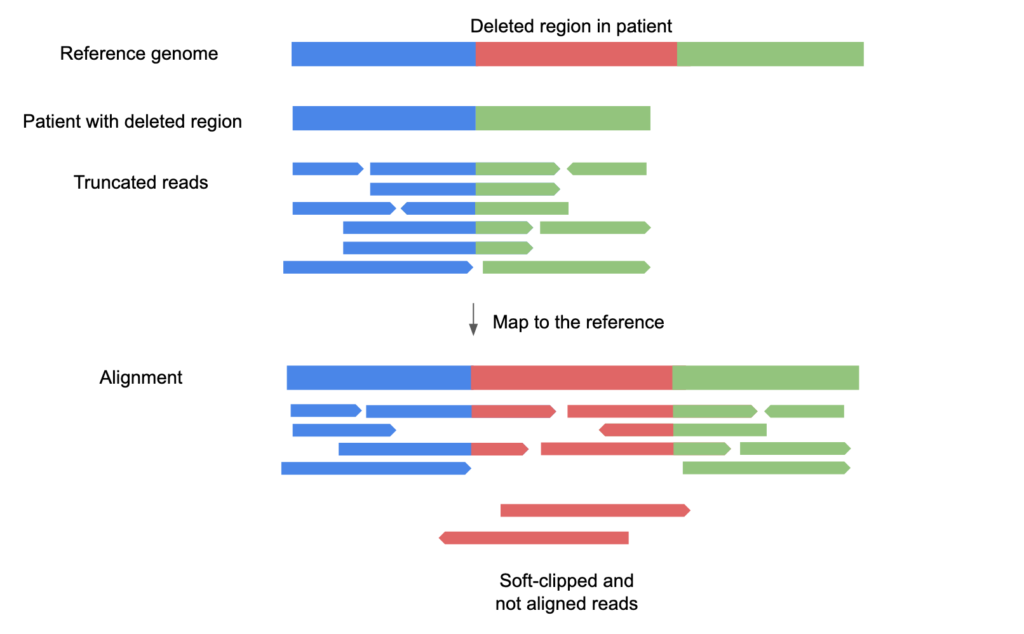
Challenges and importance of mid-sized deletion identification for genomic medicine - SeqOne Genomics
sequence alignment - What is local realignment and what is the problem it solves? - Bioinformatics Stack Exchange

Soft-clipped reads vs. improperly aligned reads at a structural variant... | Download Scientific Diagram
Sequencing artifacts derived from a library preparation method using enzymatic fragmentation | PLOS ONE
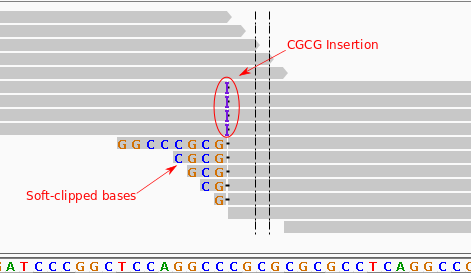
QC Fail Sequencing » Soft-clipping of reads may add potentially unwanted alignments to repetitive regions
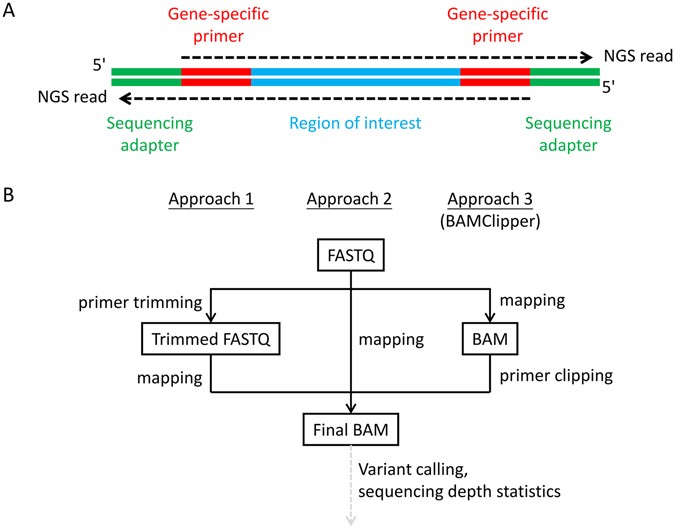
BAMClipper: removing primers from alignments to minimize false-negative mutations in amplicon next-generation sequencing | Scientific Reports
QC Fail Sequencing » Soft-clipping of reads may add potentially unwanted alignments to repetitive regions